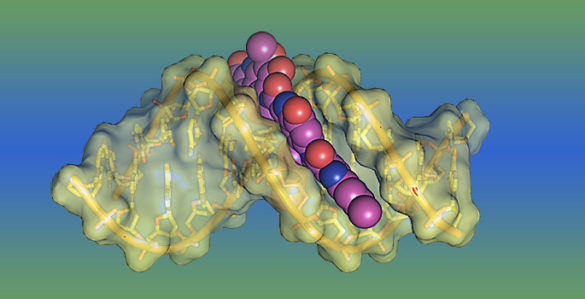
One of the most potent toxins known acts by welding the two strands of the famous double helix together in a unique fashion which foils the standard repair mechanisms cells use to protect their DNA.
A team of Vanderbilt University researchers have worked out the molecular details that explain how this bacterial toxin—yatakemycin (YTM)—prevents DNA replication. Their results, described in a paper published online July 24 by Nature Chemical Biology, explain YTM’s extraordinary toxicity and could be used to fine-tune the compound’s impressive antimicrobial and antifungal properties.
YTM is produced by some members of the Streptomyces family of soil bacteria to kill competing strains of bacteria. It belongs to a class of bacterial compounds that are currently being tested for cancer chemotherapy because their toxicity is extremely effective against tumor cells.

“In the past, we have thought about DNA repair in terms of protecting DNA against different kinds of chemical insults,” said Professor of Biological Sciences Brandt Eichman. “Now, toxins like YTM are forcing us to consider their role as part of the ongoing chemical warfare that exists among bacteria, which can have important side effects on human health.”
Cells have developed several basic types of DNA repair, including base excision repair (BER) and nucleotide excision repair (NER). BER generally fixes small lesions and NER removes large, bulky lesions.
A number of DNA toxins create bulky lesions that destabilize the double helix. However, some of the most toxic lesions bond to both strands of DNA, thereby preventing the cell’s elaborate replication machinery from separating the DNA strands so they can be copied. Normally, this distorts the DNA’s structure, which allows NER enzymes to locate the lesion and excise it.
“YTM is different,” said postdoctoral fellow Elwood Mullins. “Instead of attaching to DNA with multiple strong covalent bonds, it forms a single covalent bond and a large number of weaker, polar interactions. As a result, it stabilizes the DNA instead of destabilizing it, and it does so without distorting the DNA structure so NER enzymes can’t find it.”
“We were shocked by how much it stabilizes DNA,” Eichman added. “Normally, the DNA strands that we used in our experiments separate when they are heated to about 40 degrees [Celsius] but, with YTM added, they don’t come apart until 85 degrees.”
The Streptomyces bacteria that produce YTM have also evolved a special enzyme to protect their own DNA from the toxin. Surprisingly, this is a base excision repair enzyme—called a DNA glycosylase—that is normally limited to repairing small lesions, not the bulky adducts caused by YTM. Nevertheless, studies have shown that it is extremely effective.
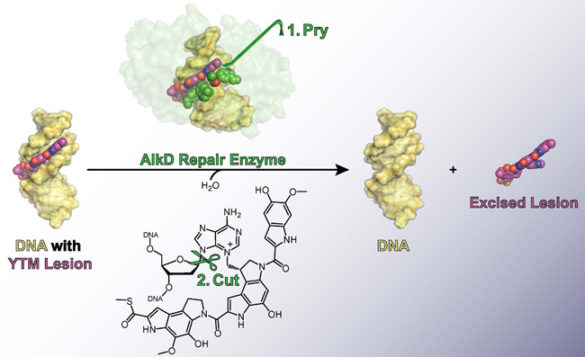
It so happens that one of Streptomyces’ competitors, Bacillus cereus, has managed to co-opt the gene that produces this particular enzyme. In Bacillus, however, the enzyme it produces—called AlkD—provides only limited protection.
In 2015, Eichman and Mullins reported that, unlike other BER enzymes, AlkD can detect and excise YTM lesions. At the time, they had no idea why it wasn’t as effective as its Streptomyces counterpart. Now they do. It turns out that AlkD tightly binds the product that it forms from a YTM lesion, inhibiting the downstream steps in the BER process that are necessary to fully return the DNA to its original, undamaged state. This drastically reduces the effectiveness of the repair process as a whole.
In recent years, biologists have discovered that animals and plants host thousands of different species of commensal bacteria and this microscopic community, called the microbiome, plays a surprisingly important role in their health and well-being. Normally, these bacteria are beneficial—for example, converting indigestible foods into digestible forms—but they can also cause problems, such as the stomach bacteria Heliobacter pylori that can cause inflammation that produces ulcers.
“We know that bacteria produce compounds like YTM when they are under stress,” Eichman observed. “The negative effects this has on their hosts is an unfortunate side effect. So it is very important that we learn as much as we can about how these bacterial toxins work and how bacteria defend against them.”
Graduate research assistant Rongxin Shi also participated in the research, which was funded by National Science Foundation grant MCB-1517695, National Institutes of Health grant R01 ES019625 and Department of Energy’s Office of Science grant DE-AC02-06CH11357.