On their way to decoding the genome of every organism in a major branch of the tree of life – that of the humble budding yeasts – a team of evolutionary biologists successfully reconstructed the genomic and metabolic characteristics of the last common ancestor of today’s more than 1,000 species of the organism.
Cornelius Vanderbilt Chair in Biological Sciences Antonis Rokas and his collaborators started by sequencing the genomes of a staggering 220 yeast species and examining them together with more than 100 others already sequenced.
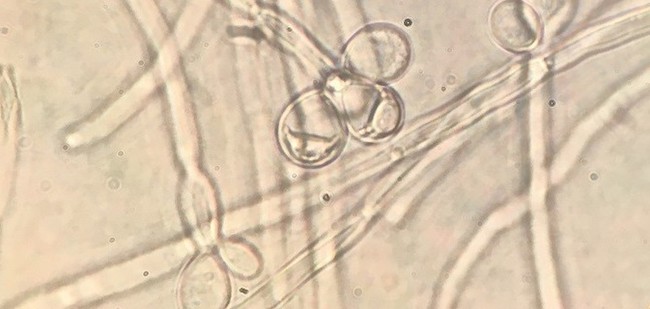
This massive sequencing effort, coupled with information of budding yeast diets, allowed them to not only retrace budding yeast evolution, but also to reconstruct the metabolic traits of their ancient parent. This last ancestor shared by all living budding yeasts lived approximately 400 million years ago and was far more metabolically diverse than the average modern budding yeast, able to live on a third more food sources than its typical modern counterparts.
It appears that, for budding yeasts, “loss is more,” Rokas says.
“One major implication of our work is that this loss of metabolic capabilities represents a major mode of budding yeast evolution,” he says. “Shedding abilities may sometimes make you better adapted to your particular environment. We know this to be true for obligate parasites, and others have theorized that this may be the case more generally, but we believe this is the first time we can actually see evidence of ‘loss is more’ for an entire lineage of free-living eukaryotic organisms.”
Rokas’ Vanderbilt team worked on this project in collaboration with Professor Chris Todd Hittinger’s team at the University of Wisconsin-Madison and with the late Clete Kurtzman of the U.S. Department of Agriculture. The team’s study, which is entitled “Tempo and mode of genome evolution in the budding yeast subphylum,” was led by Vanderbilt postdoctoral scholars Xing-Xing Shen and Xiaofan Zhou and University of Wisconsin postdoctoral scholars Dana Opulente and Jacek Kominek and appears today in the journal Cell. Jacob Steenwyk, a graduate student in the Biological Sciences graduate program, as well as former Vanderbilt postdoctoral scholars Jennifer Wisecaver and Mingshuang Wang also contributed.
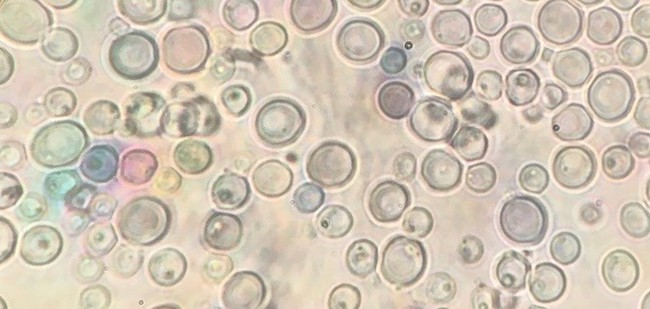
Budding yeasts are common in nearly every environment on the planet, perhaps best known for the achievements of a handful of species in the beer, wine, and bread industries or, less attractively, sending people to the drug store to treat infections. They’re known for remarkable metabolic diversity, making them of great interest to various biotechnological, food and beverage industries.
Nonetheless, the metabolic dexterity of most species in the lineage remain “untapped,” and Rokas hopes the new genomes change this.
“The Budding Yeast Common Ancestor was a metabolically complex organism,” Rokas said. “The average modern yeast can grow on about 20 substrates. Their ancestor, BYCA, could grow on 30. So that means, over the course of 400 million years of evolution, the typical species of budding yeast lost about a third of the metabolic capacity on what they were able to consume. We’ve recorded instances of this loss in several different pathways for several different food sources, such as losing the ability to utilize nitrogen as a single source.”
He said the team’s effort with budding yeasts is part of the answer to increasing calls for sequencing the genomes of all known species.
The research was funded by multiple sources, including National Science Foundation grants DEB-1442113 (Rokas), DEB-1442148 (Hittinger and Kurtzman), and a Guggenheim fellowship to Rokas