
If Guinness World Records had a category for the largest human-designed protein, then a team of Vanderbilt chemists would have just claimed it.
They have designed and successfully synthesized a variant of a protein that nature uses to manufacture the essential amino acid histidine. It is more than twice the size of the previous record holder, a protein created by researchers at the University of Washington in 2003.
The synthetic protein, designated FLR, validates a new approach which the Vanderbilt scientists have developed that allows them to design functional artificial proteins substantially larger than previously possible.
“[rquote]We now have the algorithms we need to engineer large proteins with shapes that you don’t see in nature.[/rquote] This gives us the tools we need to create new, more effective antibodies and other beneficial proteins,” said Jens Meiler, the associate professor of chemistry at Vanderbilt who led the effort.
Recently, protein engineers have verified a potential treatment strategy for HIV by using designed protein vaccines in mice and have designed artificial proteins that mimic antibodies in broadly neutralizing flu infections. The technique developed at Vanderbilt promises to expand the scope of these efforts substantially.
That is important because proteins are the most important molecules in living cells. They perform most of the vital tasks that take place within a living organism. There are hundreds of thousands of different proteins. They come in a variety of shapes and sizes. They can be round or long and thin, rigid or flexible. But they are all made out of linear chains of 20 amino acids encoded in the genome of the organism.
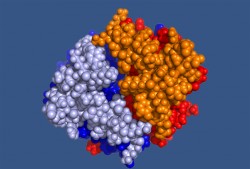

Proteins assume this variety of shapes and sizes by the manner in which they bunch and fold. This complex process takes two steps. First, small numbers of adjacent amino acids form what scientists call secondary structures: the most common of which are a rod-like spiral shape called the alpha-helix and a flat, pleated shape called the beta-sheet. These secondary structures, in turn, interact, fold and coil to form the protein’s three-dimensional shape, which is the key to its function.
Over the past 10 years an increasing number of proteins that don’t exist in nature have been designed “in silico” (in a computer). Scientists use sophisticated protein modeling software that incorporates the relevant laws of physics and chemistry to find amino acid sequences that fold into stable forms and have specific functions.
Imagine making a necklace 10 beads long with beads that come in 20 different colors. There are more than 10 trillion different combinations to choose among. This provides an idea of the complexity involved in designing novel proteins. For a protein of a given size, the modeling software creates millions of versions by putting each amino acid in every position and evaluating the stability of the resulting molecule. This takes a tremendous amount of computing power which skyrockets as the length of the protein increases.
“The current limit of this approach, even using the fastest supercomputers, is about 120 amino acids,” said Meiler. The previous record holder contained 106 amino acids. The newly designed protein contains 242 amino acids. The Vanderbilt group got around this limit by modifying the widely used protein engineering platform called ROSETTA so that it can incorporate symmetry in the design process.
Their success provides new support for a controversial theory about protein evolution called the gene duplication and fusion hypothesis. The advantage of small proteins is that they can evolve rapidly in response to changing conditions, but larger proteins can perform more complex functions. Nature found a way to get both advantages by selecting small proteins that can interact with other copies of themselves to form larger proteins, which are called dimers. Once useful dimers have been created the gene that coded for the original protein is duplicated and fused to form a new gene that can directly produce the dimer. After it is created, the dimer gene is gradually modified by natural selection to make it more efficient or develop new functions.
Because they have two identical halves, dimers have a large degree of symmetry. By taking these symmetries into account, the Vanderbilt group was able to substantially reduce the amount of computing time required to create the FLR protein. Using 400 processors of the supercomputer at Vanderbilt’s Advanced Computing Center for Research and Education, it took 10 days of continuous processing to find the most stable configuration.
To check the accuracy of their design, the researchers synthesized the DNA sequence that produces the protein, inserted it in E.coli bacteria and determined that they produced the protein and it folded properly.
The FLR protein assumes a 3-D shape called a TIM barrel, which is found in 10 percent of proteins and is particularly prevalent among enzymes. It is formed from eight beta strands that are surrounded by eight alpha helices arranged in a hexagonal shape like a tiny barrel.
The paper reporting this achievement appears in the Nov. 16 issue of the Journal of American Chemical Society and is available online. Members of Meiler’s team are research assistant Carie Fortenberry, undergraduate students Elizabeth Bowman, Will Proffitt, and Brent Dorr and research assistant professors of biochemistry Joel Harp and Laura Mizoue. The research was supported by grants from the Defense Advanced Research Projects Agency’s protein design project and the National Science Foundation.