Drug combinations used for the treatment of non-small-cell lung cancer (NSCLC) and melanoma aren’t as effective as they could be. Oncologists haven’t had the right tools to predict drug interactions, other than in costly clinical trials.
That could change with a new algorithm developed by a cross-disciplinary Vanderbilt University team for calculating drug synergy. The new approach distinguishes drug interactions which translate to quicker, more effective killing of tumors from interactions that may result in fewer side effects.
The team used it in 500,000 individual tests of drug combinations applied to live cancer cells and believe it can be applied to treating many more kinds of cancer.
“When it comes to finding new cancer drug candidates, somehow academia and the pharmaceutical industry are mixing up potency and efficacy,” said Professor of Biochemistry and Pharmacology Vito Quaranta, “That’s a fundamental flaw.”
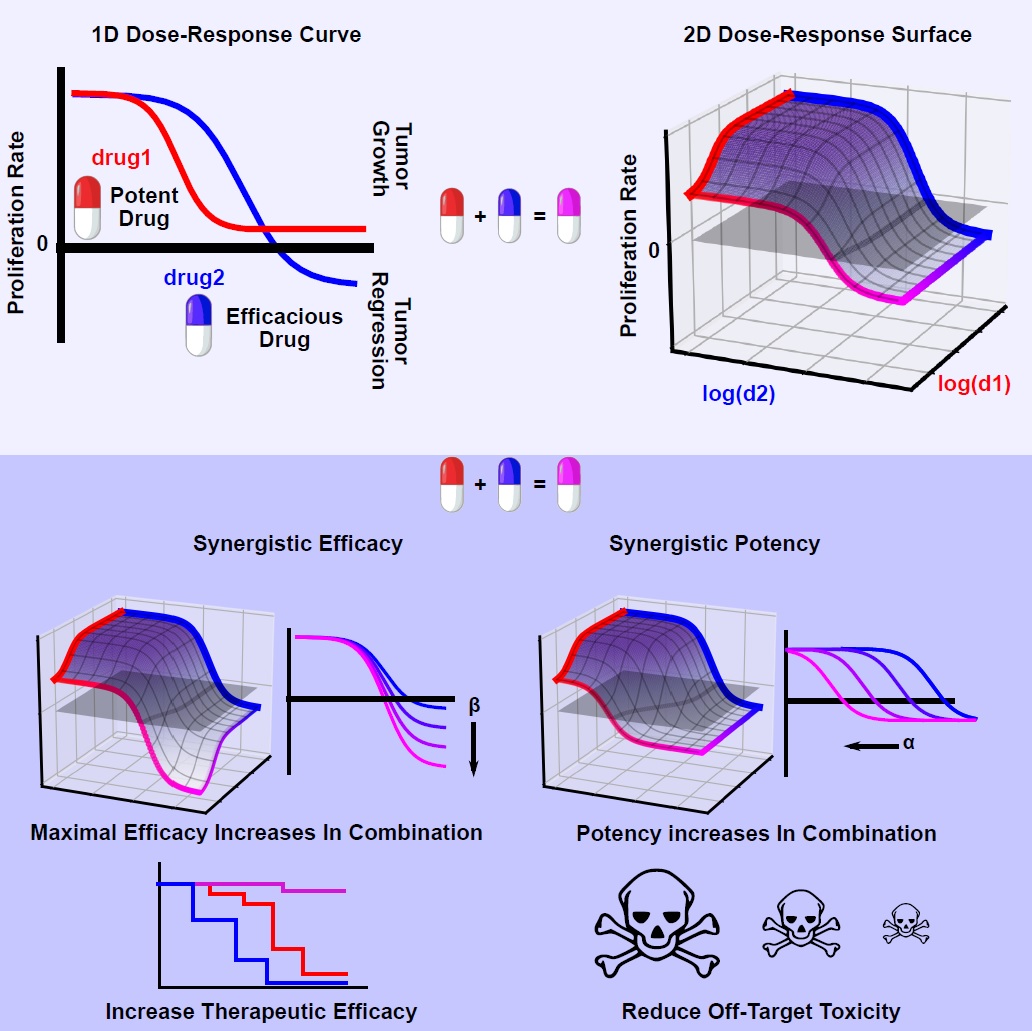
Synergy of potency is ideal to reduce drug dosage, and therefore possible side effects. Synergy of efficacy, instead, aims to raise the therapeutic ceiling, which is especially desirable in diseases prone to recur due to incomplete clearance, such as non-small cell lung cancer.
The team is calling the algorithm MuSyC, which stands for Multi-dimensional Synergy of Combinations. Synergy frameworks prior to MuSyC did not distinguish between potency and efficacy, resulting in misleading impressions of a combinations true interaction. Their work appears today in the journal Cell Systems in a paper titled “Quantifying Drug Combination Synergy Along Potency and Efficacy Axes.”
Assistant Professor of Medicine Christine Lovly and research assistant professor Darren Tyson, in conjunction with Joshua Bauer in the Vanderbilt Institute for Chemical Biology High Throughput Screening Core, conducted more than 500,000 measurements of 12,000 drug combination conditions. They sought to calculate the synergistic profile of 64 anti-cancer drugs in combination with standard of care osimertinib in NSCLC, which targets mutant EGFR.
Grouping the drugs into four categories — mitotic checkpoint, epigenetic regulators, receptors and/or channels, and kinases — revealed drug class trends obfuscated by prior frameworks. Specifically, the team found drug combinations targeting a subset of mitotic checkpoints as well as epigenetic regulators with EGFR dramatically increased the rate cancer cells were dying compared to either alone. For example, the combination of osimertinib with vindesine, a drug targeting microtubules, was 70 percent more efficacious than osimertinib alone.
“Knowing how combinations interact is crucial to increasing therapeutic efficacy and patient quality of life. Now we have a mathematical framework for quantifying and distinguishing how a combination is performing along these goals,” said Christian Meyer, graduate student in the Chemical and Physical Biology Program and first author on the paper.
The researchers foresee MuSyC will change the way drug combinations are discovered, translated, and deployed in many diseases.
Now the team is studying the interactions of three drug combinations, as well as investigating drug combinations in more complex pre-clinical models such as cancer organoids.
An early version of the work was presented to the Cancer Systems Biology Consortium in 2018, and Meyer will present again at the Keynote Symposium on Phenotypic Drug Discovery in March.
Quaranta is director of the Quantitative Systems Biology Center and principal investigator of the NCI Cancer Systems Biology Center at Vanderbilt. Information for the Vanderbilt Institute for Chemical Biology High Throughput Screening Core can be found at http://www.vanderbilt.edu/hts/
The work was supported by the National Science Foundation Graduate Student Fellowship Program Award 1445197; National Institutes of Health grants U54-CA217450, R01-186193, U01-CA215845, P30-CA086485, R01-CA121210 and P01-CA129243; National Cancer Institute R50-CA211206; Ruth L. Kirschstein National Research Service Award 2T32HL094296-06A1; V Foundation Scholar-in-Training Award; an AACR Genentech Career Development Award; a Damon Runyon Clinical Investigator Award; a LUNGevity Career Development Award; and a Lung Cancer Foundation of America/International Association for the Study of Lung Cancer Lori Monroe Scholarship.