by Carol Rouzer
The growing problem of antibiotic resistance seriously impedes the treatment of bacterial infections. Contributing to drug resistance are proteins that export toxic molecules from the bacterial cell, such as the Escherichia coli EmrE transporter.
It is known that alternate binding of a proton or a positively charged substrate molecule to a glutamic acid in the active site of EmrE is key to the transporter’s function, but the way in which this mutually exclusive binding leads to export of the substrate remains a mystery.
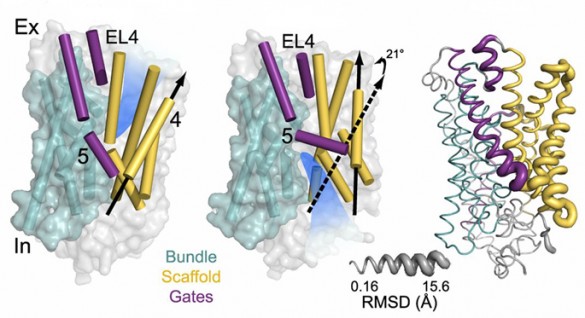
Hassane Mchaourab, Ph.D., Jens Meiler, Ph.D., and colleagues combined double electron-electron resonance spectroscopy and molecular modeling to explore EmrE dynamics. They report in Proceedings of the National Academy of Sciences that EmrE assumes different shapes when it binds a proton, a substrate, or neither.
These shape changes give EmrE the flexibility needed to bind a wide array of substrates, to prevent wasteful transport of protons, and to transport substrate in the correct direction. The results provide new insight into a key mechanism of antibacterial resistance.
The research was supported by National Institutes of Health grants GM077659 and GM080403.
Send suggestions for articles to highlight in Aliquots and any other feedback about the column to aliquots@vanderbilt.edu