Defects in kinase signaling pathways can cause cancer. Thus kinases, enzymes that regulate complex cellular processes, are among the most important cancer drug targets today.
Yet kinase inhibitors often miss the mark because the aberrant enzymes they target are part of dynamic signaling networks. Interactions with other enzymes essentially allow the “bad actors” to “escape.”
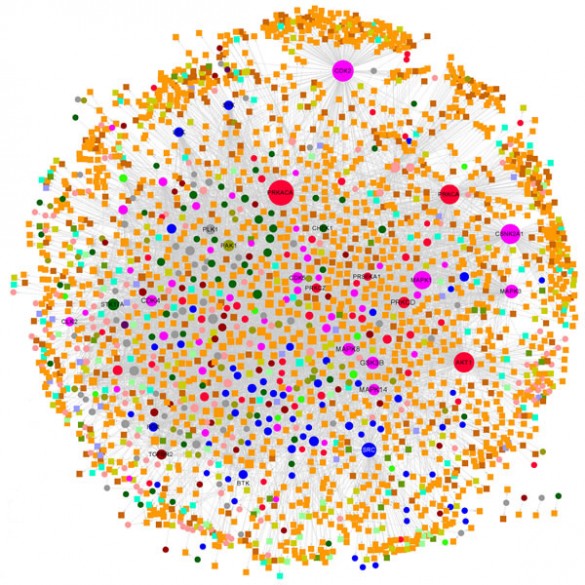
To better understand this problem, Zhongming Zhao, Ph.D., and colleagues constructed a comprehensive human kinome interactome map that incorporated three networks: interactions between the “kinome” (set of kinases) and the phosphorylation substrates the enzymes act upon, physical protein-protein interactions, and a separate set (interactome) of three-dimensional, atomic-resolution protein-protein interactions.
Reporting in the journal Oncotarget, the scientists found that distinct network hubs of kinases create a high risk of drug resistance by “feedback” or “crosstalk” mechanisms.
There is still much to be learned about the human kinome interactome. Yet “network-based modeling potentially opens a new avenue for … rational kinase inhibitor design in individualized cancer therapy,” they concluded.
The study was supported in part by National Institutes of Health grants LM011177, CA167695, CA095103, CA098131 and CA068485.
Send suggestions for articles to highlight in Aliquots and any other feedback about the column to aliquots@vanderbilt.edu















